Show summary Hide summary
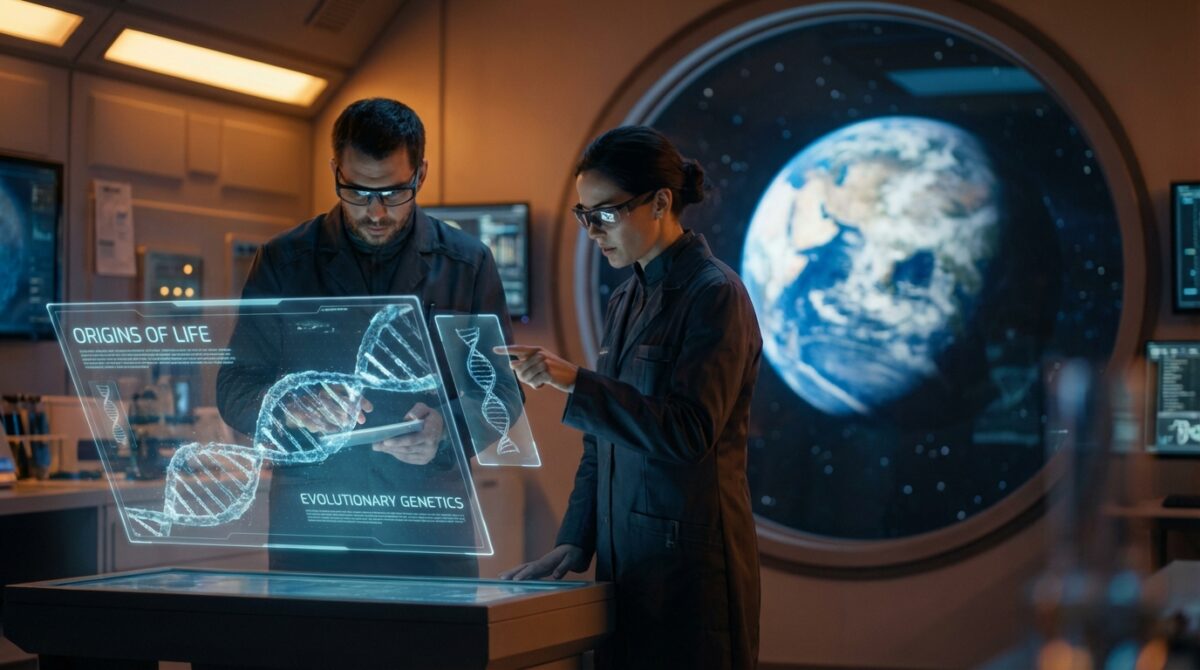
Some of the oldest genetic traces on Earth may be older than all known life itself. New work in evolutionary genetics suggests that a handful of ancient genes were already duplicated and working long before the first shared ancestor of every organism alive today ever appeared.
This changes what scientists can realistically ask about the origins of life on Earth. Instead of stopping at the “last universal common ancestor” (LUCA), researchers now have a data-driven way to peek into an even earlier, more mysterious phase of evolution.
Ancient genes predating life as we know it
The study, published in Cell Genomics, comes from a team led by Aaron Goldman (Oberlin College), working with Greg Fournier at MIT and Betül Kaçar at the University of Wisconsin–Madison. They examined what we now know is the deepest accessible layer of genetic history: universal paralogs.
The epic return of epicyon: prehistoric america’s fearsome bone-crushing predator
How the Rethinking Economics Movement Is Transforming Education
These genes seem to have duplicated before LUCA, roughly four billion years ago. That means they preserve traces of cellular biology from lineages that no longer exist, a kind of prehistoric black box recording from the dawn of life. Like the DNA study that recast the history of the Beachy Head Woman, this project shows how much hidden history can sit inside a few key sequences of DNA.
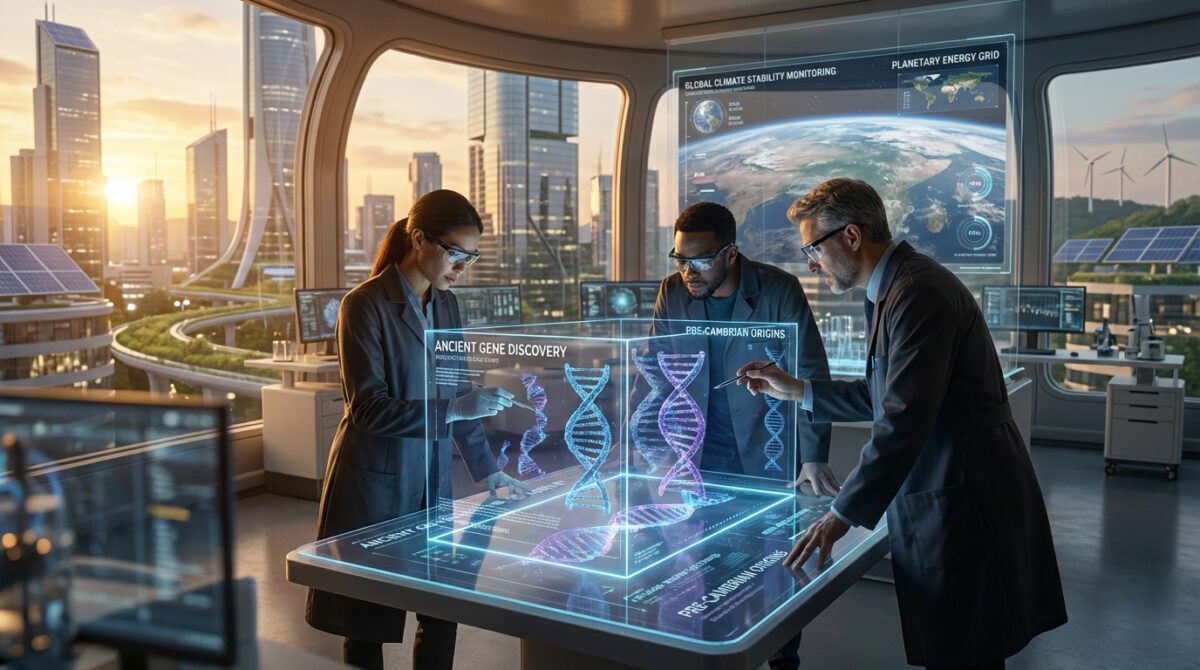

How scientists traced these prehistoric genetic signals
To track these ancient genes, the team used one core method: compare gene families that appear in at least two copies in almost every known organism, then reconstruct their evolutionary tree backward in time. If a gene and its “twin” are everywhere, the duplication probably happened before LUCA, not after.
Modern AI-assisted sequence analysis and powerful computing clusters made it possible to sift through vast genomic databases. The researchers then applied established evolutionary models to estimate when specific duplications occurred and how the corresponding proteins might have looked and behaved billions of years ago.
What universal paralogs reveal about the first cells
Out of all the genes checked, only a small, rare set qualified as true universal paralogs. Every one of them fell into just two jobs: building proteins or moving molecules across cell membranes. Nothing else made the cut.
This pattern matters. It suggests that even before LUCA, early cells were already investing heavily in two operations that still dominate biology today: turning genetic code into working proteins, and regulating what enters or leaves a cell. That gives a more concrete picture of what the earliest functional cells were doing, rather than treating them as abstract “primitive blobs.”
Reconstructing an ancestral membrane machine
To go beyond patterns and into performance, Goldman’s group focused on one universal paralog family involved in inserting proteins into cell membranes. Using evolutionary reconstruction, they inferred the sequence of the original ancestral protein, then synthesized and tested it in the lab.
The reconstructed protein was simpler than its modern descendants, yet it still attached to membranes and interacted with the machinery that makes proteins. It likely acted as a minimal “insertion helper,” guiding early enzymes into rudimentary membranes. That single result turns a speculative story about early life into something closer to a replayed experiment.
Key findings that reshape the story of life on Earth
Several central insights emerge from this work, with important caveats about what can and cannot be claimed as cause-and-effect. The study does not prove how life started, but it narrows the range of plausible scenarios for the first cells capable of sustained evolution.
- Universal paralogs predate LUCA: Their shared duplication pattern strongly suggests an origin before the last universal common ancestor, pushing accessible genetic history beyond four billion years.
- Early biology focused on proteins and membranes: All known universal paralogs are tied to protein synthesis or membrane transport, hinting that these were among the earliest stable cellular functions.
- Reconstructed ancestral proteins still work: Lab tests show that simplified ancient versions can perform core tasks, supporting the idea that early life ran on lean, efficient molecular systems.
- Genetic “fossils” complement geological evidence: These gene families add a molecular layer to the same deep-time window that geologists probe in ancient rocks and isotopes.
Like ambitious microbiome research that tries to unlock the power of our microbiome, this project shows how far scientists can push inference from indirect data, while still grounding claims in measurable sequences and testable reconstructions.
What the numbers and uncertainties tell us
The scope is vast but not infinite: the team worked with all currently known universal paralog families, a set numbered in just the dozens, out of millions of gene families cataloged across genomes. Confidence in their deep ancestry comes from converging phylogenetic trees and the near-global distribution of these genes in bacteria, archaea, and eukaryotes.
However, precise dates carry broad confidence intervals, because processes that old leave only faint statistical signals. The authors hedge timing estimates carefully, treating them as ranges rather than fixed points. Correlation between universal paralog functions and early cellular needs is strong, yet causation remains an inference, not a direct measurement.
How this changes the search for life’s origins
For researchers like Leila, a young evolutionary biologist building her first project on early genetics, this framework reshapes strategy. Instead of guessing which genes might be ancient, she can target documented universal paralogs and their reconstructed ancestors, then design experiments around membrane behavior and protein assembly.
Policy makers and funding agencies also gain a clearer map. Projects that connect AI-driven genomic mining with wet-lab reconstructions now sit at the frontier of prebiotic science, in the same way that sea turtle resilience studies have updated climate expectations in work like new insights on global warming and marine life. The common thread is careful use of models to extend limited data, without treating those models as oracles.
Limits, open questions and future directions
Several constraints keep the picture incomplete. Universal paralogs are rare, so they offer only a partial catalogue of prehistoric cell biology. Many early genes may have been lost entirely, leaving no trace in modern genomes. Horizontal gene transfer and gene loss over billions of years also blur some of the signals that phylogenetic trees rely on.
Yet those gaps define the next phase of research. As genomic sequencing covers more obscure microbes, and AI tools refine ancestral reconstructions, new candidate universal paralogs may emerge. Each one could add a small but solid piece to the jigsaw puzzle of prehistoric cellular life on Earth, moving speculation into the realm of testable science.
What is the last universal common ancestor (LUCA)?
LUCA is the most recent organism from which all current life on Earth descends. It lived roughly four billion years ago and already used DNA, proteins, and cell membranes. It is not the very first life-form, but the earliest one that current evolutionary methods can reliably study.
What are universal paralogs in genetics?
Universal paralogs are gene families that exist in at least two copies in nearly all known organisms. Their widespread duplication pattern suggests that the original gene split happened before LUCA, making them valuable markers of very early cellular evolution.
Do these ancient genes explain how life began?
They do not fully explain how life started, and they do not prove a specific origin scenario. Instead, they highlight which functions—such as protein production and membrane transport—were already active very early, helping to constrain and refine origin-of-life models.
How do scientists reconstruct prehistoric proteins?
Researchers compare related gene sequences across many species, build an evolutionary tree and infer the most likely ancestral sequence at key branching points. They then synthesize this ancient-like sequence in the lab and test the resulting protein’s behavior under controlled conditions.
Could similar methods help search for life beyond Earth?
Life’s Origins Sticky Theory: Goo Sparks Existence
Toxic Metals Discovered in Bananas Post Brazil Mining
Indirectly, yes. By clarifying which genetic and cellular functions appear robust across deep time, scientists can better guess what kinds of biochemistry might evolve elsewhere. However, without actual extraterrestrial samples, this remains informed speculation rather than direct evidence.