Show summary Hide summary
- Ancient plant DNA that refuses to disappear
- How researchers hunted invisible genetic switches
- Three evolution rules shaping plant gene regulation
- A regulatory atlas for future plant biology
- From lab insights to climate-ready crops
- What is a conserved non-coding sequence in plants?
- How old are the genetic switches discovered in this study?
- Why were plant regulatory sequences harder to find than in animals?
- How can this discovery help improve crops?
- What tools did researchers use to identify these sequences?
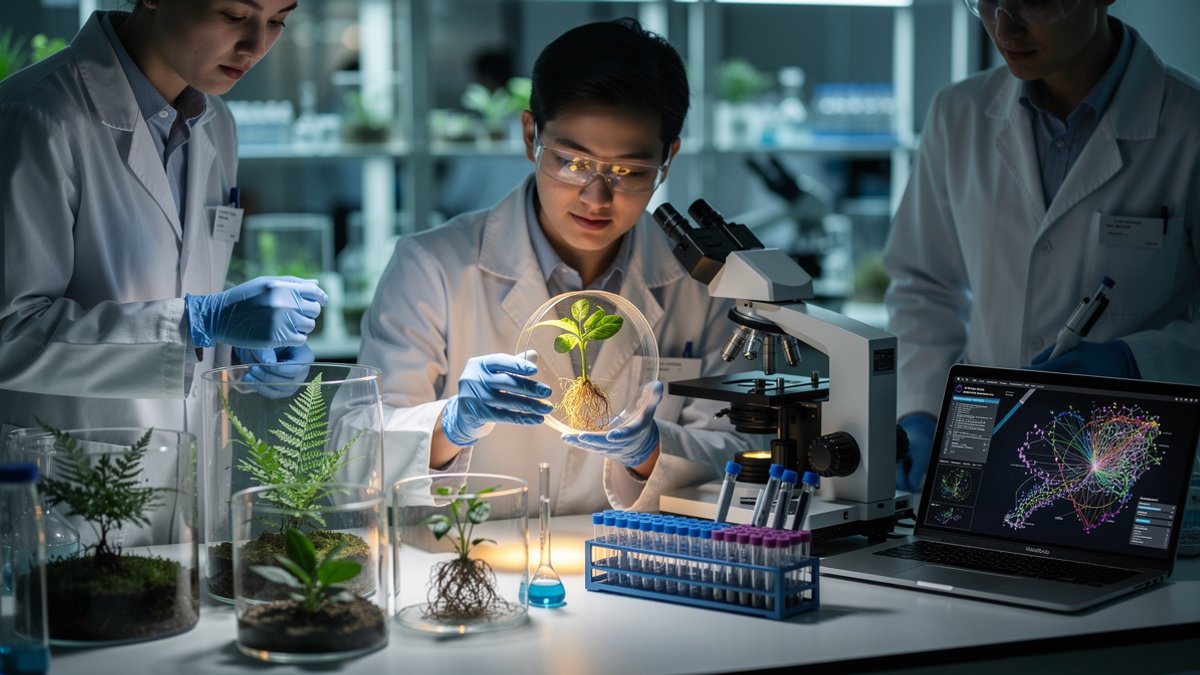
Hidden inside leaves and roots, Genetic Switches older than the first dinosaurs are still guiding how modern Plants grow and adapt. These fragments of Ancient DNA have survived more than 400 Million Years of evolution and are now rewriting what you know about Plant Biology and Gene Regulation.
Imagine following a single regulatory sequence from the earliest land plants to today’s crops. That is what an international team of Researchers has effectively done, using cutting-edge Genetics and Molecular Biology tools to map how these hidden switches shape life on Earth.
Ancient plant DNA that refuses to disappear
The new work centers on pieces of non-coding DNA that do not build proteins but act as control panels for genes. These regions, called conserved non-coding sequences (CNSs), function like dimmers and timers for key developmental genes. They tell a gene when to activate, in which tissue, and at what intensity.
Researchers Unlock 3D Printing Technique for One of Earth’s Toughest Metals
Scientists Uncover Hedgehogs’ Ability to Hear Ultrasound, Offering New Hope to Avoid Road Accidents
Using a custom-built algorithm named Conservatory, teams from Cold Spring Harbor Laboratory and partner institutes scanned 314 plant genomes from 284 species. They uncovered more than 2.3 million CNSs that stayed recognizable despite hundreds of millions of years of divergence between mosses, trees and flowering crops.
From pre-flowering ancestors to modern crops
The most striking result concerns timing. Some CNSs appear to predate the split between flowering and non-flowering plants, a split that goes back more than 400 Million Years. That means identical regulatory motifs helped control early land plants long before seeds or petals evolved.
This level of conservation rivals stories told in human biology, such as the project that mapped 7 million human cells across organs to decode aging. In plants, the same logic applies: track persistent patterns in old DNA, and you reveal the long-term rules of life’s design.
How researchers hunted invisible genetic switches
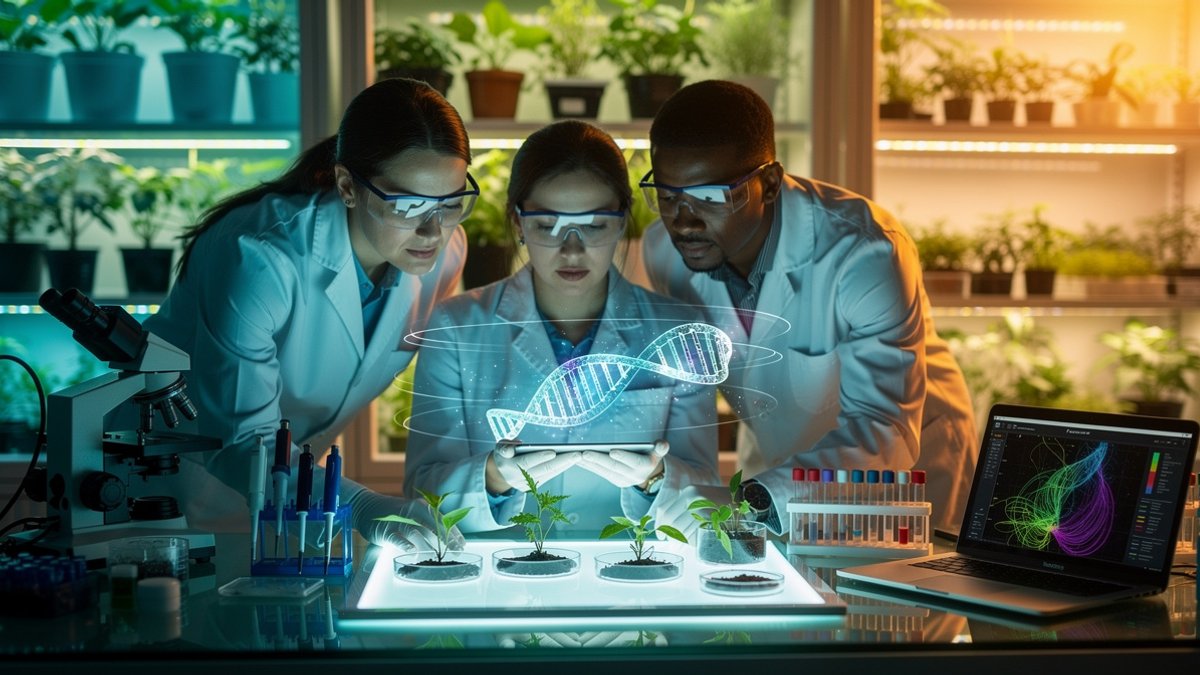

Classic genome comparisons usually align long DNA segments and look for matching blocks. For plants, that approach kept failing. Genomes have swollen, shrunk and reshuffled through repeated duplications and rearrangements, scrambling long-distance alignments and masking short regulatory elements.
The Conservatory project took a different angle. The team focused on tiny neighborhoods of genes, examining how small gene clusters are arranged and conserved. By tracking local order rather than long stretches, they could spot CNSs that moved or stretched but kept their relative positions and functions. Read more about harnessing quantum mysteries in genome research.
Editing DNA to test what really matters
Finding patterns in silico was not enough. To prove these CNSs are true Genetic Switches, the scientists used genome editing to remove or alter specific sequences. Plants with edited CNSs showed disrupted development, confirming that these old fragments still drive growth and organ formation.
This experimental strategy echoes other frontier work in Molecular Biology, like studies that map DNA architecture before embryos even form. In all these cases, targeted edits reveal which elements behave like master controls rather than genomic noise.
Three evolution rules shaping plant gene regulation
By following CNSs across species and time, the researchers distilled three recurring patterns that describe how regulatory DNA evolves in plants. These rules help explain both long-term stability and innovation inside plant genomes.
First, the physical distance between CNSs and genes may change, but their order on the chromosome tends to remain. Second, when plant genomes are rearranged, CNSs can become connected to new genes, rewiring regulatory networks without inventing completely new sequences. Third, after gene duplication, ancestral CNSs often persist alongside duplicate genes, supplying raw material for new regulatory variants.
Old switches as engines of novelty
These three rules together sketch a dynamic picture. Ancient switches do not just endure; they are repurposed. A CNS originally tuning leaf shape can, after a genomic reshuffle, start influencing flower timing or root growth while still keeping its core sequence features.
That reuse parallels medical work where genetic switches keep organs healthy in humans. In both kingdoms, evolution often modifies existing regulators rather than inventing code from scratch. Old modules get redeployed into new developmental contexts.
A regulatory atlas for future plant biology
The Conservatory dataset effectively forms an atlas of regulatory conservation across land plants. It covers dozens of major crops, including cereals, legumes and vegetables, as well as their wild relatives that endure heat, drought or poor soils. For plant scientists, it now acts like GPS for hidden regulatory zones.
A breeding program, for example, can overlay field performance data with CNS maps. If a wild relative tolerates arid conditions, breeders can pinpoint conserved switches near stress-response genes and test whether fine-tuning those elements boosts resilience in an elite line without harming yield.
From lab insights to climate-ready crops
In a world already reshaped by climate change, this regulatory atlas comes at a timely moment. Similar to how new studies suggest sea turtles may cope with warming better than feared, plants also carry deep evolutionary tools for adaptation locked in DNA.
By targeting conserved regulatory sequences instead of whole genes, crop engineers can pursue softer adjustments: shifting flowering dates, optimizing root depth or modulating water use. The long survival of these switches across Million Years signals that they offer robust levers rather than unstable hacks.
- Decoding history: CNSs reveal how regulatory networks guided plant evolution from early land colonizers to intensive agriculture.
- Guiding breeding: Maps of conserved switches help select varieties with better stress responses without sacrificing taste or yield.
- Informing engineering: Targeted edits in CNSs allow refined control of Gene Regulation instead of blunt genetic changes.
- Connecting kingdoms: Parallels with animal studies highlight shared principles of genome organization and regulation.
What is a conserved non-coding sequence in plants?
A conserved non-coding sequence (CNS) is a stretch of DNA that does not code for proteins but has stayed similar across many plant species over long evolutionary periods. These regions usually act as regulatory switches, controlling when and where nearby genes are activated.
How old are the genetic switches discovered in this study?
Analyses of plant family trees indicate that some CNSs originated before flowering plants diverged from non-flowering ancestors, more than 400 million years ago. These ancient sequences have been maintained because they perform vital regulatory roles that support plant survival and development.
Why were plant regulatory sequences harder to find than in animals?
Plant genomes have undergone frequent duplications and rearrangements, which scramble long stretches of DNA and hide short regulatory motifs. Traditional alignment methods failed to follow these elements across species. The new approach, focusing on local gene neighborhoods, made it possible to detect CNSs despite large-scale genome reshuffling.
How can this discovery help improve crops?
The Urgent Quest to Crack the Greatest Challenge in Quantum Computing
Researchers Develop the Most Challenging AI Benchmark Yet—Unexpected Outcomes Emerge
By mapping where conserved regulatory DNA sits near important genes in crops and their wild relatives, breeders and engineers can fine-tune traits such as stress tolerance, flowering time, or yield stability. Editing or selecting variations in CNSs offers a precise way to adjust gene activity without changing the underlying protein-coding regions.
What tools did researchers use to identify these sequences?
The team developed a computational tool called Conservatory, which compares small gene clusters across hundreds of plant genomes to detect conserved regulatory DNA. They then validated candidate CNSs with genetic editing experiments, showing that disrupting these regions often alters plant growth and organ formation.