Show summary Hide summary
- Light-guided evolution reinventing protein engineering
- Rewiring the yeast cell cycle for molecular switching
- New color channels and smarter light sensors
- Proteins acting as single-molecule biocomputers
- Key advantages of light-guided protein design
- FAQ
- How does light-guided protein evolution differ from traditional directed evolution?
- What are the main applications of light-guided protein evolution?
- What role does optogenetics play in this approach?
- Can light-guided protein evolution improve the precision of synthetic biological systems?
- Why is yeast commonly used in experiments involving light-guided protein evolution?
Imagine proteins inside yeast cells behaving like tiny computers, flipping states on precise schedules, all directed by pulses of light. That is the promise of light-guided protein evolution: using lasers instead of pipettes to sculpt living molecular machines for switching, sensing, and computing.
Light-guided evolution reinventing protein engineering
Evolution has always been nature’s way to perform protein engineering. Random DNA changes appear, and cells carrying the most functional proteins outgrow the rest. Farmers once exploited this logic in fields and barns; modern labs now compress it into flasks and incubators.
Traditional directed evolution usually rewards proteins that stay “on” all the time. Many cellular components, however, act as molecular switching devices or logic gates, turning on briefly, then shutting off again. When experiments only prize constant activity, those subtle timing tricks decay, and proteins lose their ability to flip states reliably.
Marine Animal Virus Linked to Unusual Eye Conditions in Humans
Exploring the Powerful Impact of the Heart-Brain Connection on Your Well-being
Optovolution: when optogenetics meets natural selection
Researchers at EPFL introduced an approach called optovolution, a form of light-guided protein evolution that steers whole populations of cells using optogenetics. Instead of manually testing thousands of variants, they allow survival to depend on how a protein responds to prescribed light patterns.
In this scheme, light pulses become the fitness test. Cells hosting proteins that follow the right timing pattern divide again; the others stall or die. As reports such as this detailed overview explain, the process mirrors natural selection, but with a programmable spotlight replacing harsh, random environments.
Rewiring the yeast cell cycle for molecular switching
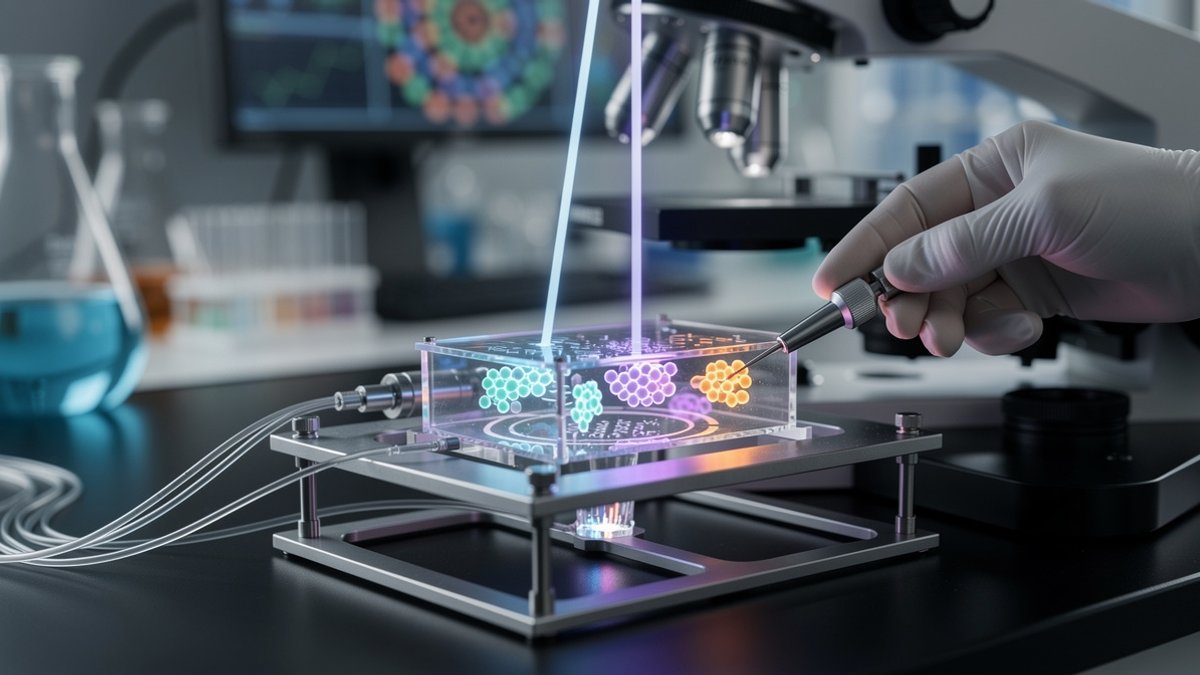

The team worked with budding yeast, a champion of both brewing and basic research. They rewired the yeast cell cycle so that division depended on the behavior of a chosen protein. That protein had to switch cleanly between active and inactive states at specific phases.
They linked the protein’s output to a cell-cycle regulator that is vital at one stage yet toxic at another. If the protein stayed “on” too long, the toxic phase killed the cell. If it switched off too early, the vital step failed. Only cells with correctly timed molecular switching continued growing, automatically enriching precise dynamic responses.
Using photocontrol as an evolutionary metronome
To orchestrate this timing, the researchers relied on photocontrol. Using light-sensitive domains fused to the protein of interest, they turned activity on or off with defined light pulses. Each yeast cycle of about ninety minutes provided a quick pass-or-fail decision.
Because selection unfolded continuously, the system did not require laborious plate screening. The rhythm of the light schedule shaped the evolving population, giving protein design a temporal dimension that classical methods rarely achieve. This metronome-like control is what pushes optovolution beyond earlier protein design strategies.
New color channels and smarter light sensors
Once the evolutionary engine was running, the team first targeted a widely used light-controlled transcription factor. Through repeated cycles, they obtained 19 new variants with sharper behavior: some responded to weaker illumination, others reduced “leakiness” in darkness, and several shifted sensitivity from blue to green.
Shifting proteins towards warmer colors has long challenged optogenetics, because pigment chemistry usually favors blue absorption. Optovolution found routes that conventional design had struggled to identify, a result echoed in analyses like this exploration of how light accelerates protein evolution. Expanded color sensitivity opens the door to multi-channel biosensing where different wavelengths independently control distinct pathways.
Evolution hacking cellular logistics for red light tools
The researchers then turned to a red-light system that initially required an added chemical cofactor. Over successive generations, an unexpected mutation knocked out a standard yeast transport protein. That single change allowed the optogenetic module to harness light-sensitive molecules already present inside the cell.
This twist matters for synthetic biology practitioners who want simpler, plug-and-play constructs. Eliminating external cofactors reduces experimental overhead and makes red-light tools more attractive for long-term cultures, bioreactors, or complex tissue models that cannot be dosed continuously.
Proteins acting as single-molecule biocomputers
Optovolution did not stop at light sensors. The EPFL group also evolved a transcription factor that performed an AND logic operation, an entry point to biocomputing. This protein activated gene expression only when it simultaneously received a light cue and a chemical signal.
Such a design echoes digital electronics: a gate fires only when multiple conditions are met. Inside cells, this type of decision-making could coordinate drug release, control differentiation, or filter noisy stimuli. Reports on proteins that switch like tiny computers show how rapidly this logic-based view of biology is spreading.
From lab demo to future cellular circuits
Consider a fictional startup, LumoCell, aiming to program immune cells that attack tumors only when they detect a cancer metabolite and a surgeon’s light cue. With an optovolution workflow, LumoCell could evolve receptors and transcription factors that implement exactly that two-input logic, tested automatically over many generations.
Step by step, such platforms could yield layered circuits, with several AND, OR, and NOT gates built from functional proteins. The result would be living systems whose internal state machines rival simple electronic controllers, but run on enzymes and photons instead of silicon and electrons.
Key advantages of light-guided protein design
For researchers or engineers planning the next generation of cellular tools, this light-guided protein evolution framework brings multiple advantages that go beyond classical mutagenesis and screening.
- Temporal precision: proteins are selected for correct timing, not just strength, enabling finely tuned molecular switching.
- Complex behaviors: multi-state responses and logic-gate functions evolve continuously, ideal for advanced biosensing and biocomputing.
- Automation-ready: survival-based selection cuts down on manual assays and scales naturally in fermenters or microfluidic devices.
- Spectral diversity: new color channels, from blue to green to red, support multiplexed optogenetics in crowded cellular environments.
Together, these features reposition directed evolution as a tool for dynamic control rather than static optimization, aligning lab-built systems with the rhythms of real cellular life.
What does light-guided evolution change compared to classic directed evolution?
Light-guided protein evolution, or optovolution, introduces time-dependent selection based on specific light patterns. Proteins are rewarded not only for being active, but for switching on and off at the right moments. This enables the evolution of dynamic, multi-state behaviors such as precise molecular switching and logic-like decisions inside cells.
Why use yeast for evolving functional proteins with light?
Yeast divides quickly, is genetically tractable, and tolerates extensive engineering of its cell cycle. By wiring a target protein to an essential cell-cycle regulator, researchers can make yeast survival depend on the protein’s dynamic behavior. This creates a robust living platform to evolve light-responsive and computation-capable proteins.
How does optovolution support biosensing applications?
By evolving proteins that respond to multiple inputs, such as different light colors or combined light and chemicals, optovolution yields highly selective biosensing modules. These sensors can be integrated into synthetic biology circuits to detect environmental cues, disease markers, or metabolic states and trigger tailored downstream responses.
Can this approach be extended beyond optogenetics tools?
Yes. Although many demonstrations involve light-sensitive domains, the same logic can evolve transcription factors, receptors, or enzymes whose activity controls cell survival in a time-dependent manner. Any protein whose output can be coupled to a growth decision can, in principle, be optimized with light-guided protein evolution schemes.
What are the main future uses in biocomputing?
Future biocomputing applications include living logic circuits that decide when to release therapies, regulate metabolism in bioreactors, or coordinate complex developmental programs. By evolving proteins that implement AND, OR, and other logic operations using light and chemical cues, optovolution brings programmable protein-based computation closer to practical deployment.
FAQ
How does light-guided protein evolution differ from traditional directed evolution?
Light-guided protein evolution uses precise light patterns instead of constant conditions to select for proteins with specific switching behaviours. This allows researchers to evolve proteins for dynamic tasks like sensing or timing, rather than just always-on activity.
What are the main applications of light-guided protein evolution?
This technique is especially useful for engineering proteins that function as switches, logic gates, or sensors within cells. It opens possibilities for biological computing and better control in synthetic biology.
What role does optogenetics play in this approach?
Optogenetics enables scientists to control protein behaviour in living cells with light. In light-guided protein evolution, optogenetics helps steer cell populations by rewarding only those that respond correctly to light cues.
Can light-guided protein evolution improve the precision of synthetic biological systems?
Which Type of Olive Oil Boosts Brain Health the Most?
Scientists Confirm: E-Cigarettes Outperform Patches and Gum as Smoking Cessation Tools
Yes, by evolving proteins to respond predictably to light, researchers can design biological systems with more accurate timing and switching. This is important for applications requiring tight control or programmable behaviours.
Why is yeast commonly used in experiments involving light-guided protein evolution?
Yeast is easy to manipulate genetically and grows quickly, making it ideal for evolutionary experiments. It also shares many features with higher cells, allowing discoveries to be translated to more complex systems.